Product Description
CD1E is a member of the CD1 family of transmembrane glycoproteins, which are structurally related to the major histocompatibility complex (MHC) proteins and form heterodimers with beta-2-microglobulin. The CD1 proteins mediate the presentation of primarily lipid and glycolipid antigens of self or microbial origin to T cells. The human genome contains five CD1 family genes organized in a cluster on chromosome 1. The CD1 family members are thought to differ in their cellular localization and specificity for particular lipid ligands. CD1E localizes within Golgi compartments, endosomes, and lysosomes, and is cleaved into a stable soluble form. The soluble form is required for the intracellular processing of some glycolipids into a form that can be presented by other CD1 family members. Recombinant human CD1E protein, fused to His-tag at N-terminus, was expressed in E.coli.
Biovision | 7308 | CD1E human recombinant DataSheet
Biomolecule/Target: CD1E
Synonyms: T-cell surface glycoprotein CD1e, CD1A, R2
Alternates names: T-cell surface glycoprotein CD1e, CD1A, R2
Taglines: Mediates the presentation of primarily lipid and glycolipid antigens of self or microbial origin to T cells
NCBI Gene ID #: N/A
NCBI Gene Symbol: N/A
Gene Source: Human
Accession #: N/A
Recombinant: Yes
Source: E. coli
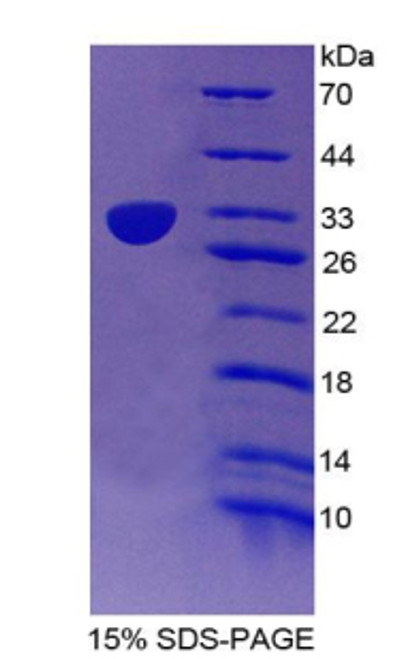
Purity by SDS-PAGEs: 99%
Assay: SDS-PAGE
Purity: N/A
Assay #2: N/A
Endotoxin Level: < 1.0 EU per 1 microgram of protein
Activity (Specifications/test method): N/A
Biological activity: N/A
Results: N/A
Binding Capacity: N/A
Unit Definition: N/A
Molecular Weight: 33.1 kDa (297 aa, 32-305 aa + His tag), confirmed by MALDI-TOF.
Concentration: 1 mg/ml
Appearance: Liquid
Physical form description: 1 mg/ml in 20 mM Tris-HCl buffer (pH 8.0) containing 10% glycerol, 0.4M Urea
Reconstitution Instructions: N/A
Amino acid sequence: MGSSHHHHHH SSGLVPRGSH MGSEEQLSFR MLQTSSFANH SWAHSEGSGW LGDLQTHGWD TVLGTIRFLK PWSHGNFSKQ ELKNLQSLFQ LYFHSFIQIV QASAGQFQLE YPFEIQILAG CRMNAPQIFL NMAYQGSDFL SFQGISWEPS PGAGIRAQNI CKVLNRYLDI KEILQSLLGH TCPRFLAGLM EAGESELKRK VKPEAWLSCG PSPGPGRLQL VCHVSGFYPK PVWVMWMRGE QEQRGTQRGD VLPNADETWY LRATLDVAAG EAAGLSCRVK HSSLGGHDLI IHWGGYS
 Euro
Euro
 USD
USD
 British Pound
British Pound
 NULL
NULL